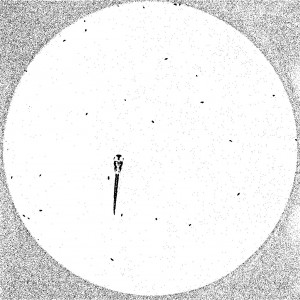
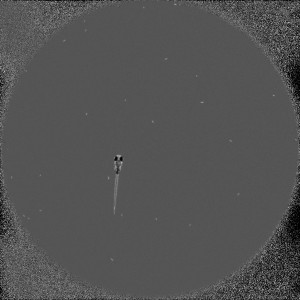
To quantitatively describe the ability of prey capture in zebrafish larvae, I need to count the number of paramecia consumed by the larva. Here is a frame from the time-lapse recording of the prey capture behavior. In a typical assay, I put one zebrafish larva and about 30 paramecia in a recording chamber and record for 11 minutes at 10 fps (exposure time: 5msec). The larva will eat most of the paramecia during recording. This prey capture behavior can be used as an assay to test how an experimental manipulation affects the prey capturing ability.
How can I count the paramecia in ImageJ? First, I obtain a background image by making an averaged image. To do this, call Image > Stacks > Z Project… > Average Intensity.
 (Note that embedded image is in JPEG just for the sake of reducing the file size on this blog).
(Note that embedded image is in JPEG just for the sake of reducing the file size on this blog).
Now I divide the movie by this background image. Process > Image Calculator… > Divide (check both “Create new window” and “32-bit (float) result”). Whether we need background subtraction or not depends on the image. In some cases, background subtraction does not give a better result.
Image > Adjust > Brightness/Contrast… > Auto, then Image > Type > 8-bit . Now you can clearly see the paramecia in a uniform background.
Image > Adjuts > Threshold… > Upper Threshold Level:94, Apply.
Edit > Invert (Process all images)
Now I set a oval ROI on the circular chamber and analyze particles. Analyze > Analyze Particles… > size (pixel^2): 18-72, Circularity:0.50-1.00, Show:Masks, Check “Display results”, “Clear results”, “Summarize”, “Exclude on edges”, and click OK (Process all images).
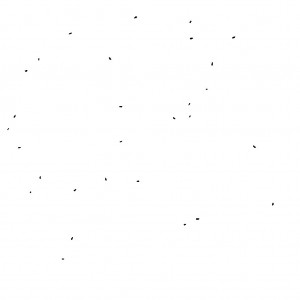
As a result, the paramecia are extracted. This is the “Mask of ..” image (Different frame from the above).
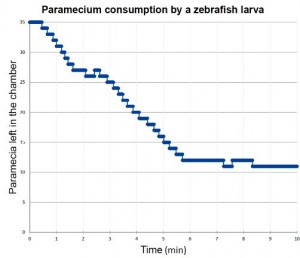
The text window named “Summary of ..” gives the “Count”. The numbers fluctuates among frames because some paramecia may be missed because they were near the edge of the chamber or there happened to be some noises in each frame. Because I don’t need exact paramecium count frame-by-frame, I average for 1-min interval (moving average for 600 frame time bin on Excel). What I need in this experiment is to know how many paramecia are consumed over minutes. A typical result looks like this (after averaging). On average, this zebrafish larva captured one paramecium in every 15 seconds.
Automating the paramecium counting
Particle analysis is a slow process (depending on how many particles are present in the image). In my case, one file takes about 30 min. So, it will take 15 hours to process 30 recordings. It is exhausting if I need to do it manually. Spending some time on writing an ImageJ macro can save a huge amount of time and energy.
The size of a recording file is 6.4GB ( 1024 x 1024 pixels x 6600 frames). My desktop PC has 32GB memory, not enough memory to do division of the 6.4GB images. So I have to work on a half of the movie at one time. I wrote an ImageJ macro (which worked in ImageJ 1.49v on my data files on Windows7).
This macro will:
- obtain file names in a specified folder and works on each AVI file.
- It opens one AVI file,
- makes an averaged image,
- duplicates the 1st and 2nd half of the movie
- and closes the original movie,
- does image division to subtract background
- and saves the resulted images for the future use (in RAW fromat),
- thresholds to make binary images,
- particle analyzes,
- and saves the result as a text file.
- Then it moves on to the next AVI file in the folder
- and repeats processing until all are done.
Note: I found that a large (>6GB) stack could not be saved properly in AVI or TIFF format. Only RAW format worked in my macro for unknown reason.
By the way, you do not need to write a macro from scratch. Macro recording is available. Plugins > Macros > Record… This does not do everything that I want but it helps a lot.